Virtual Library
Start Your Search
Joseph B Shrager
Author of
-
+
P2.05 - Interventional Diagnostic/Pulmonology (ID 168)
- Event: WCLC 2019
- Type: Poster Viewing in the Exhibit Hall
- Track: Interventional Diagnostics/Pulmonology
- Presentations: 1
- Moderators:
- Coordinates: 9/09/2019, 10:15 - 18:15, Exhibit Hall
-
+
P2.05-01 - Broad Genomic Profiling of Bronchoalveolar Lavage Fluid in Lung Cancer (ID 615)
10:15 - 18:15 | Author(s): Joseph B Shrager
- Abstract
Background
We hypothesized that tumor-derived mutations from non-small cell lung cancer (NSCLC) are readily detectable in bronchoalveolar lavage (BAL). To explore our hypothesis, we employed the CAncer Personalized Profiling by deep Sequencing (CAPP-Seq) method to identify somatic mutations in BAL compared to blood.
Method
We profiled 200 matching lavage, plasma, and PBMC samples from a total of 38 NSCLC patients and 21 controls. We first applied a tumor-informed calling approach to most sensitively detect mutations in BAL and plasma. We then applied a tumor-naïve mutation calling strategy to explore the effect of field cancerization in at risk patients with lung nodules or who smoked. Last, we developed a BAL mutation classifier to differentiate patients with cancer from those without and compared the performance of this classifier to BAL cytology.
Result
Tumors were primarily lung adenocarcinomas (84%) and mostly early stage disease (I-II 71%; III-IV 29%). We called a median of 4 mutations per tumor. TP53 and KRAS were the most frequently detected variants in tumor (47% and 35% respectively) and lavage cell free (cf) DNA (38% and 26% respectively). Using a tumor-informed approach, we detected significantly more variants in lavage cfDNA than in plasma from cancer patients (p<0.001) and variants were more frequently called in lavage cfDNA than in plasma from cancer patients (77% vs. 41%, p=0.004). As expected, tumor-naïve calling resulted in fewer variants detected in both sample types when compared to tumor-informed calling but we identified more tumor mutations (p<0.001) and more putative cancer driver mutations (p< 0.003) in lavage cfDNA than in plasma. Mutations of cancer driver genes at the patient level and average %VAF at the gene level were significantly lower in lavage cfDNA controls compared to cancer patients (p=0.017 and p=0.016 respectively). Since we also detected mutations in controls, presumably secondary to field cancerization and somatic mosaicism, we developed a risk score of mutation features to classify whether a BAL specimen was likely to come from a cancer patient or control. At a risk score level that identified all non-cancers as benign (A), this BAL classifier of 11 mutation features identified more cancers than cytology for all stages of lung cancer (65% vs 12%, p=0.001) and in stage I/II disease (50% vs. 20%, p=0.25; B).
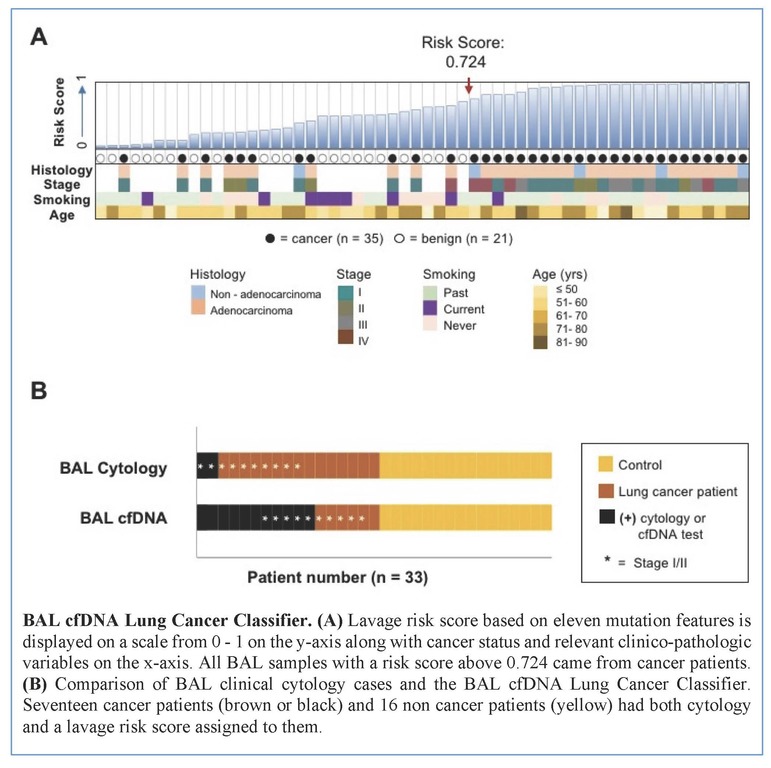
Conclusion
We show here that BAL genomic profiling may augment plasma profiling and BAL cytology for diagnosing and profiling NSCLC. Validation studies will be required to confirm our findings.


