Virtual Library
Start Your Search
Kaettipong Kamprerasart
Author of
-
+
P2.01 - Advanced NSCLC (Not CME Accredited Session) (ID 950)
- Event: WCLC 2018
- Type: Poster Viewing in the Exhibit Hall
- Track:
- Presentations: 1
- Moderators:
- Coordinates: 9/25/2018, 16:45 - 18:00, Exhibit Hall
-
+
P2.01-90 - PD-L1 Expression as a Predictive Biomarker in Advanced Non-Small Cell Lung Cancer Patients with or without EGFR Mutation (ID 13528)
16:45 - 18:00 | Author(s): Kaettipong Kamprerasart
- Abstract
Background
The prognostic value of PD-L1 expression and its clinical relevance of NSCLC is controversy. The impact of PD-L1 expression as the predictive biomarker for EGFR-TKIs treatment is needed to explore.
a9ded1e5ce5d75814730bb4caaf49419 Method
Medical records of metastatic NSCLC during September 2015-2016 had been reviewed. PD-L1 immunohistochemistry (IHC) staining with Antibody clone 22C3 was used. The PD-L1 positive was defined by tumor proportion score (TPS) > 1%.
4c3880bb027f159e801041b1021e88e8 Result
204 patients were included. Patients with positive PD-L1 expression had significantly increased numbers of metastatic sites (P=0.009) and lung metastasis (P=0.045) compared to PD-L1 negative patients. Overall survival (OS) was longer in PD-L1 negative patients (22.7 months) compared to PD-L1 positive groups (13.6 months) (HR=1.48; P=0.03). Median OS were significantly different with the number of 7.2, 11.1, 25.7, 42.6 months in EGFR-/PD-L1+, EGFR-/PD-L1-, EGFR+/PD-L1+ and EGFR+/PD-L1- , respectively (P<0.001). Among EGFR positive patients, mOS of T790M-/PD-L1+, T790M-/PD-L1-, T790M+/PD-L1-, and T790M+/PD-L1+ were 22.1, 28, 42.6 and 48.4 months, respectively (P=0.03).
Patient characteristics categorized by PD-L1 expression (N = 204) Characteristics PD-L1 Negative
N=134 (65.69%)
PD-L1 Positive
N=70 (34.31%)
P value Age
- Median age (range)
- < 65 years
- > 65 years
65 (36-85)
64 (47.76)
70 (52.24)
65 (35-86)
33 (47.14
37 (52.86)
0.933 Sex
- Male
- Female
62 (46.27)
72 )53.73)
37 (52.86)
33 (47.14)
0.371 ECOG PS
- 0-1
- > 2
116 (86.57)
18 (13.43)
57 (82.61
12 (17.39)
0.452 Smoking status
- Never
- Ex-smoker
- Current smoker
82 (61.65)
36 (27.07)
15 (11.28)
36 (52.17)
18 (26.09)
15 (21.74)
0.131 Mean smoking pack-year (range) 29.72 (2-100) 25.68 (2-40) 0.873 Initial staging
- Recurrent
- Denovo metastasis
32 (23.88)
102 (76.12)
14 (20)
56 (80)
0.529 Histology
- Adenocarcinoma
- Squamous cell carcinoma
- Adenosquamous carcinoma
- Others
117 (87.31)
1 (0.75)
2 (1.49)
14 (10.45)
58 (84.06)
3 (4.35)
2 (2.9)
6 (8.7)
0.288 EGFR mutation
- Negative
- Positive
47 (35.07)
87 (64.93)
32 (45.71)
38 (54.29)
0.092 Exon 19 deletion
- No
- Yes
87 (64.93)
47 (35.07)
50 (71.43)
20 (28.57)
0.348 L858R
- No
- Yes
100 (74.63)
34 (25.37)
53 (75.71)
17 (24.29)
0.865 ALK results
- Negative
- Positive
79 (92.94)
6 (7.06)
47 (97.92)
1 (2.08)
0.421 Number of site of metastasis
- 0-1
- > 2
88 (65.67)
46 (34.33)
37 (52.86)
33 (47.14)
0.009 Lung metastasis
- No
- Yes
92 (69.17)
41 (30.83)
38 (54.29)
32 (45.71)
0.045 Bone metastasis
- No
- Yes
101 (75.37)
33 (24.63)
53 (75.71)
17 (24.29)
0.957 Liver metastasis
- No
- Yes
122 (91.04)
12 (8.96)
62 (88.57)
8 (11.43)
0.573 Pleural metastasis
- No
- Yes
91 (67.91)
43 (32.09)
62 (88.57)
20 (28.57)
0.606 Brain metastasis
- No
- Yes
117 (87.31)
17 (12.69)
58 (82.86)
12 (17.14)
0.387 Adrenal metastasis
- No
- Yes
122 (91.04)
12 (8.96)
62 (88.57)
8 (11.43)
0.573
8eea62084ca7e541d918e823422bd82e Conclusion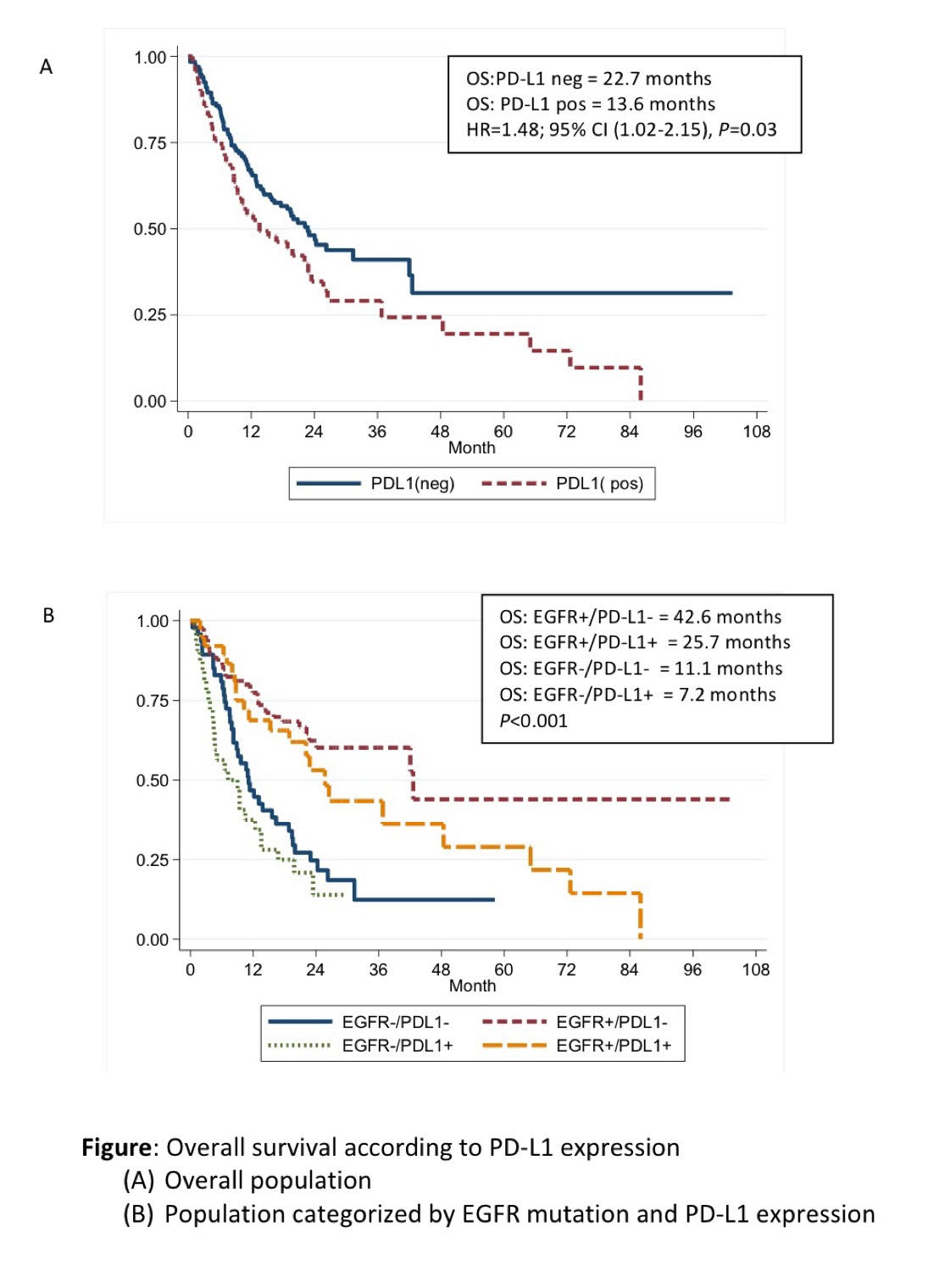
PD-L1 expression was associated with poorer survival outcomes among advanced NSCLC patients regardless of EGFR mutation status. PD-L1 expression is also the potential of predictive biomarker for EGFR TKIs treatment. The larger studies are needed to identify the prognostic and predictive values in T790M mutation population.
6f8b794f3246b0c1e1780bb4d4d5dc53
-
+
P3.09 - Pathology (Not CME Accredited Session) (ID 975)
- Event: WCLC 2018
- Type: Poster Viewing in the Exhibit Hall
- Track:
- Presentations: 1
- Moderators:
- Coordinates: 9/26/2018, 12:00 - 13:30, Exhibit Hall
-
+
P3.09-08 - Tumor Heterogeneity and Molecular Profile of NSCLC in Thai Population (ID 14016)
12:00 - 13:30 | Author(s): Kaettipong Kamprerasart
- Abstract
Background
Oncogenic driven mutation is the key to develop targeted therapy in lung cancer. Different ethnicity and tumor heterogeneity affect the prevalence of molecular alteration. This study aimed to explore the unique molecular profile of lung adenocarcinoma in Thai population.
a9ded1e5ce5d75814730bb4caaf49419 Method
We studied 166 lung adenocarcinoma patients’ molecular profile using Next Generation Sequencing (NGS) on 45 genes lung cancer panel (Ion Torrent system). Variants from NGS with coverage of higher than 1000X and cut off at 2% alternate variant frequency were considered positive. We validated the positive mutation of EGFR, BRAF, and KRAS by Real- Time PCR using the Amoy DX kit.
4c3880bb027f159e801041b1021e88e8 Result
This study found 68%(113/166) of EGFR mutation, 9.6%(16/166) of BRAF V600E, 32.5% (54/166) of KRAS mutation, 9%(15/166) of MET exon14 splice site, 4.8% (8/166) AKT mutation (E17K), 2.4% (4/166) of ROS1 mutation, 0.6% (1/166) of PIK3CA mutation (H1047R), and 0.6% (1/166) of PTEN mutation. Furthermore, we also found 40 patients (24.1%) who had more than one mutation in each person. We further validated the positive results by Real-Time PCR. Thirteen patients were obtained tissue from different organs and some with different period of time. T790M usually develop later in EGFR-positive patients who failed 1st or 2nd generation EGFR-TKI. Two patients (patient 5&9) who had lung surgery different lobe in same operation, had different mutation in tissues and one patient (patient 13) who obtained tissue from lung and pleural effusion cell block in different period of time had totally different mutation (Table1).
Table1 Tumor heterogeneity profile in lung cancer patients
case
Hetrogenety in different organ
date obtained tissue
Gene mutation
EGFR
KRAS
ROS
PTEN
AKT
MET
BRAF
1
RLL lobectomy
12-Jun-2012
Exon 19 Deletion
negative
negative
negative
negative
negative
negative
Right lung
4-May-2016
Exon 19 Deletion T790M
negative
negative
negative
negative
negative
negative
2
lymph node
16-Mar-2017
negative
negative
negative
negative
negative
negative
negative
Bone
9-Apr-2017
negative
negative
negative
negative
negative
negative
negative
3
Right upper lung biopsy
20-Jan-2016
Exon 19 Deletion
negative
negative
negative
negative
negative
negative
Lung biopsy
31-Mar-2017
T790M
negative
negative
negative
E17K
negative
negative
4
lymph node
13-Jan-2016
negative
G12A
negative
negative
negative
negative
negative
skin
17-Jan-2016
negative
G12V G12D
negative
negative
negative
negative
negative
left humerous
30-Jun-2016
negative
G12V
negative
negative
negative
negative
negative
5
RML lobectomy
13-Nov-2013
Exon 19 Deletion
G13D
negative
negative
negative
negative
negative
LUL lobectomy
14-May-2014
negative
G12D G13D
negative
R233*
negative
negative
negative
LUL lobectomy
14-May-2014
negative
G12V G12D
D2033N
negative
negative
negative
negative
6
Right pleura biopsy
11-Jan-2016
Exon 19 Deletion
negative
negative
negative
negative
negative
negative
RUL biopsy
1-Mar-2017
Exon 19 Deletion
negative
negative
negative
negative
negative
negative
7
RUL biopsy
28-Sep-2015
L858R
negative
negative
negative
negative
negative
negative
Left pleural fluid cell block
27-Jun-2016
negative
negative
negative
negative
negative
negative
negative
8
Right pleural fluid cell block
25-Mar-2016
Exon 19 Deletion
negative
negative
negative
negative
negative
negative
Left pleural fluid cell block
9-Dec-2016
T790M
negative
negative
negative
negative
negative
negative
9
RML lobectomy
14-Mar-2013
negative
G13S
negative
negative
E17K
negative
negative
RLL wedge resectionRLL lo
30-Mar-2017
G719A L861Q
negative
negative
negative
negative
negative
negative
10
RLL lobectomy
26-Aug-2013
negative
G12C G12D
negative
negative
negative
negative
negative
Left lingular lobe segmental resection
9-Oct-2014
negative
G12D
negative
negative
negative
negative
negative
LUL wedge resection
11-Feb-2016
negative
G12D
negative
negative
negative
negative
negative
LUL lobectomy
4-Jun-2017
negative
G12D
negative
negative
negative
negative
negative
LLL resection
9-Oct-2014
negative
G12D
negative
negative
negative
negative
negative
11
Right pleural cell block
21-Jul-2014
L858R
negative
negative
negative
negative
c.3028G>A exon 14 splicing
negative
Ascites
9-Jun-2017
T790M L858R
negative
negative
negative
negative
negative
negative
12
LUL biopsy
24-Feb-2015
L858R
negative
negative
negative
negative
negative
V600E
Lt lung biopsy
8-Jul-2016
L858R
negative
negative
negative
negative
negative
negative
13
RLL biopsy
14-Oct-2015
negative
negative
negative
negative
negative
c.3028+1G>T exon 14 splicing
negative
pleural fluid cell block
15-Nov-2016
Exon 19 Deletion T790M
negative
negative
negative
negative
negative
negative
8eea62084ca7e541d918e823422bd82e Conclusion
Thai populations have unique molecular alteration compared to the other ethnicities, especially, higher of BRAF V600E and MET exon14 splice site. Our population also has high co-mutation prevalence. Tumor heterogeneity is needed to explore in the larger cohort.
6f8b794f3246b0c1e1780bb4d4d5dc53


